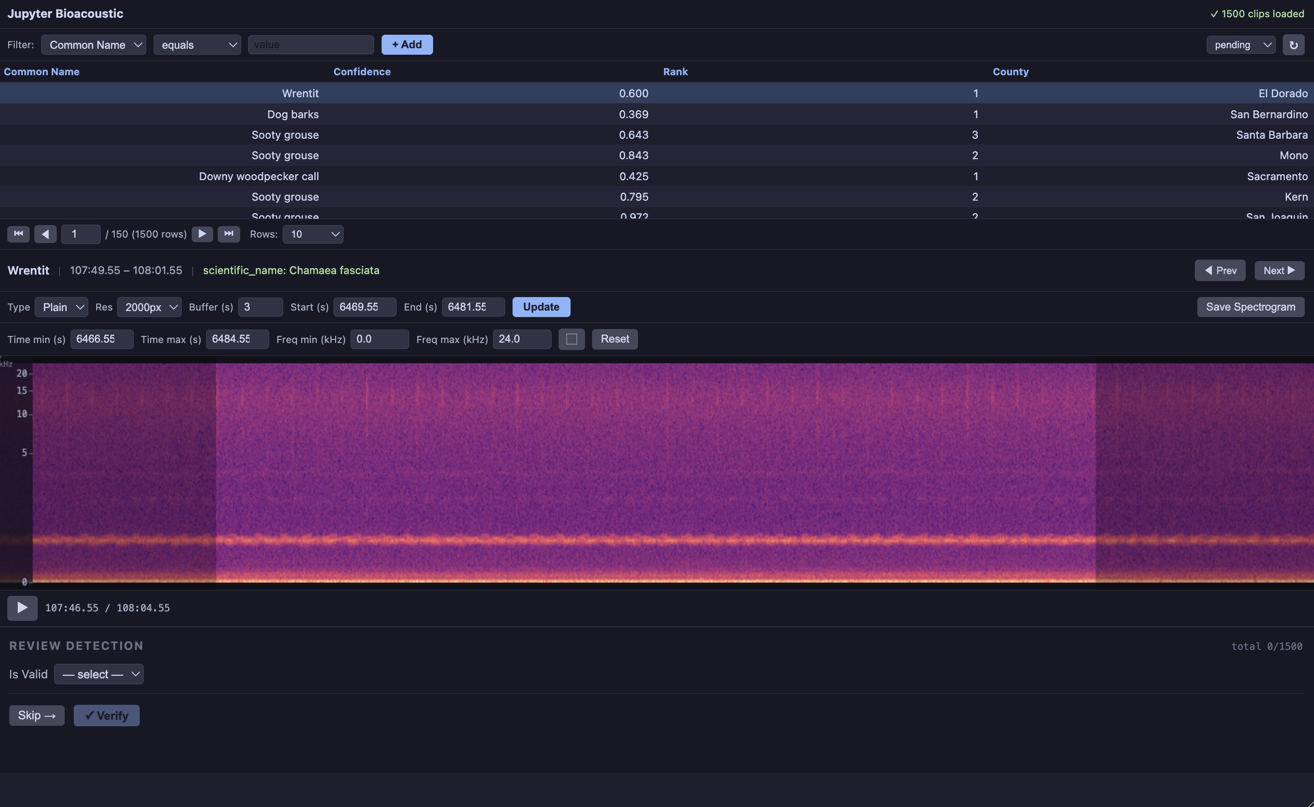
BioacousticAnnotator allows the user to configure almost every aspect of the app. Here we focus on the most minimal configurations for: a player, data collection, and model review. Each configuration below passes paramters directly to BioacousticAnnotator. Using a config file, rather than passing parameters directly will help produce cleaner, more reproducible workflows. Section 3 shows basic configuration files for the examples in section 2. See the Bioacoustic Annotators Examples Notebook for a more advanced/realistic configurations.
Author: Brookie Guzder-Williams (bguzder-williams@berkeley.edu)
Affiliation: The Eric and Wendy Schmidt Center for Data Science & Environment
Website: https://dse.berkeley.edu/
License: BSD 3-ClauseFor configuration options and advanced usage see the documentation and the geo-analysis example.
from jupyter_bioacoustic import BioacousticAnnotator
DATA = 'data/annotate-data.csv'
AUDIO = 'https://dse-soundhub.s3.us-west-2.amazonaws.com/public/audio/dev/20230522_200000.flac'
CATEGORIES = 'data/categories-small.csv'1. Player / Visualizar¶
Browse audio clips and view spectrograms — no form, no data collection. Useful for exploration and quality checks.
BioacousticAnnotator(data=DATA, audio=AUDIO).open()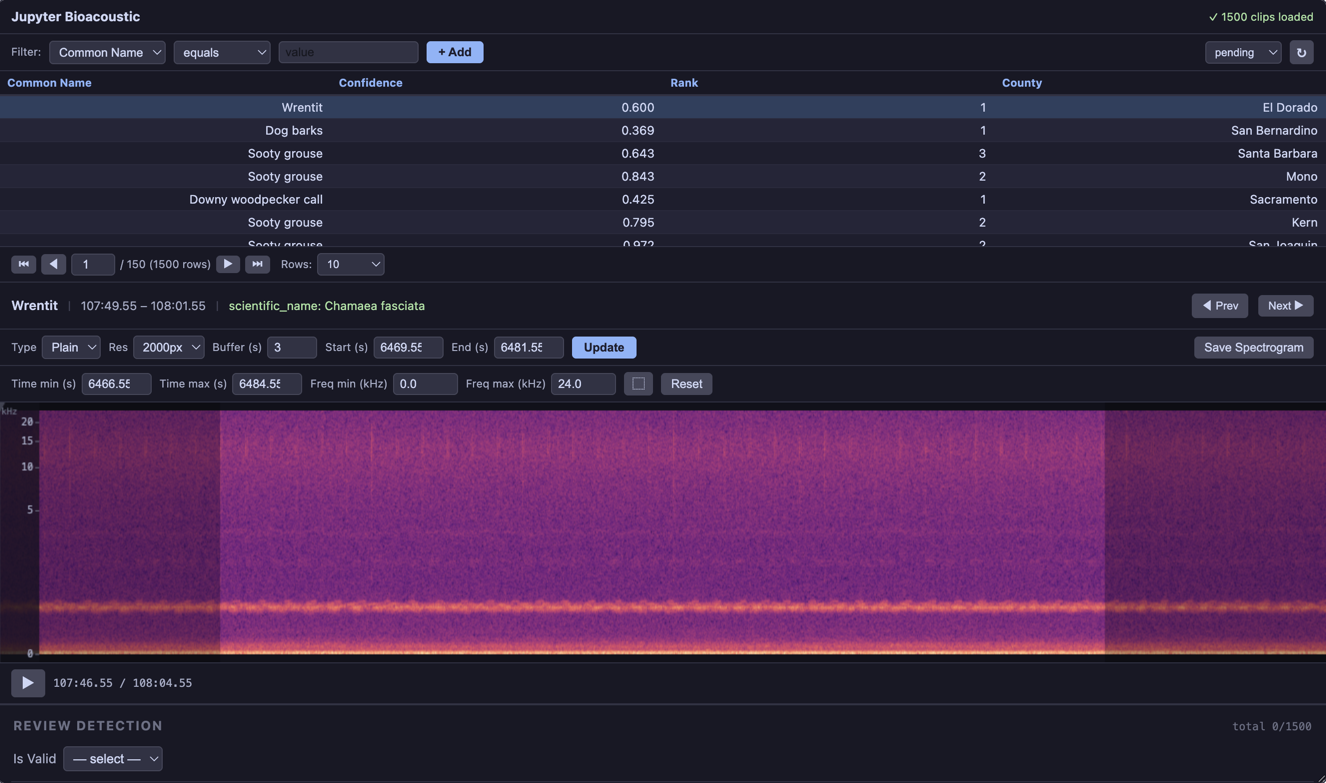
2. Simple Forms¶
Here we show two simple forms created by passing a dictionary to the form_config:
Data Collection: a simple form requiring the user to select a species for the clip
Model Review: the user is first asked if the model prediction is valid. if they select yes, the user may submit their response. If the user selects no, they must select what speicies is contained in the clip.
Note: form_config also accepts a path to yaml file. form_config may also be passed directly in a config-file (see bioacoustic
2.a Data Collection¶
Our only form element in this example is a select dropdown. This example uses the following parameters:
label: display label; used as column name if column is omittedcolumn: column written to output file (defaults to label)required: if true, submit button is disabled until value is setitems: theitemselements can take several formats, lists of strings or dicts for example. in this case we specifiy a path to a CSV and the column in the CSV to popluate the list with.
BioacousticAnnotator(
data=DATA,
audio=AUDIO,
form_config={
'select': {
'label': 'species',
'column': 'common_name',
'required': True,
'items': {
'path': CATEGORIES,
'value': 'common_name',
},
},
},
).open()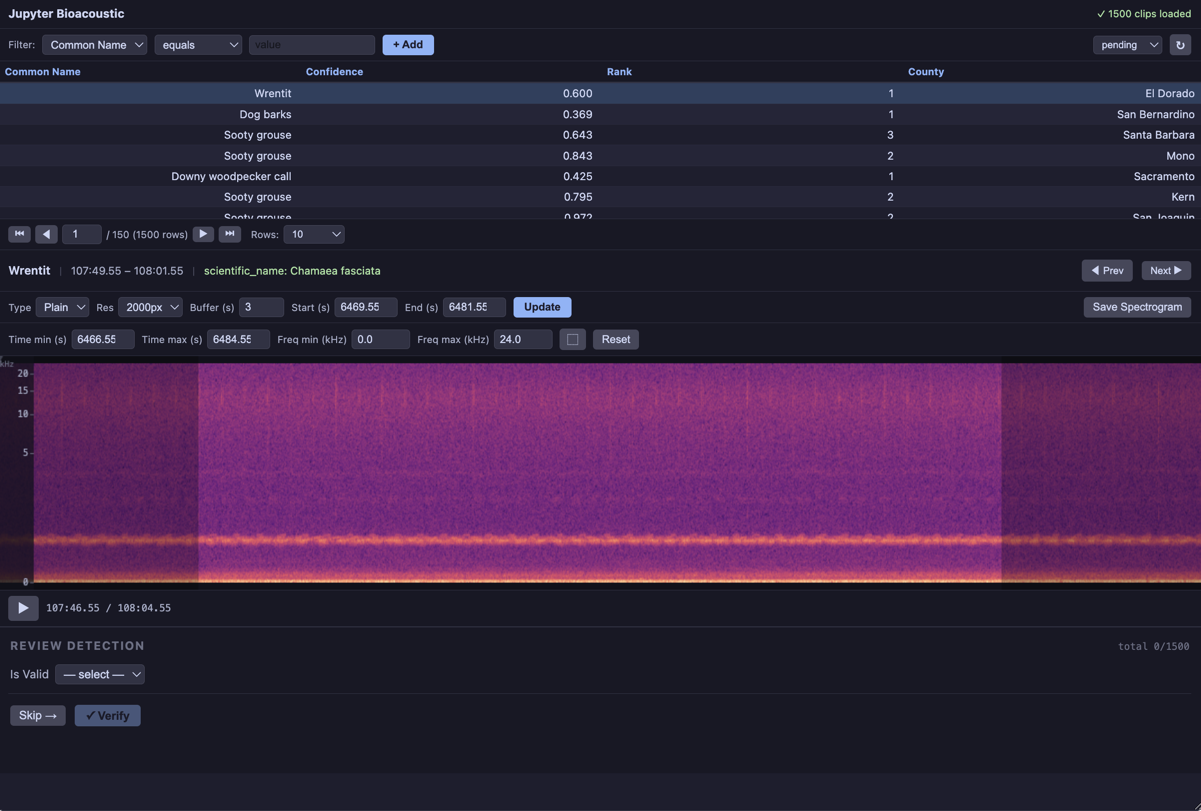
2.b Model Review¶
The form in this example in includes two elements: a “is valid” select-box, and an additional form (containing a single select-box) If the user chooses that the model has incorrectly identified the species.
Note that the correction_form select-box is similar to above with some additional elements under select.items. Namely:
filter_box:(bool)if set to true an additional textbox appears next to the dropdown that allows the user to filter through options. This can be extremely useful when there are 100s of speicies to choose fromnot_available:(bool|str)if truthy adds an additonal “not_available” value (in this case “Unknown Species”) to the list
BioacousticAnnotator(
data=DATA,
audio=AUDIO,
form_config={
'select': {
'label': 'Is Valid',
'column': 'is_valid',
'required': True,
'items': [
{ 'value': 'yes'},
{ 'value': 'no', 'form': 'correction_form'},
],
},
'correction_form': [
{'select': {
'label': 'corrected species',
'column': 'corrected_common_name',
'items': {
'path': CATEGORIES,
'value': 'common_name',
'required': True,
'filter_box': True,
'not_available': 'Unknown Species',
},
}},
],
},
).open()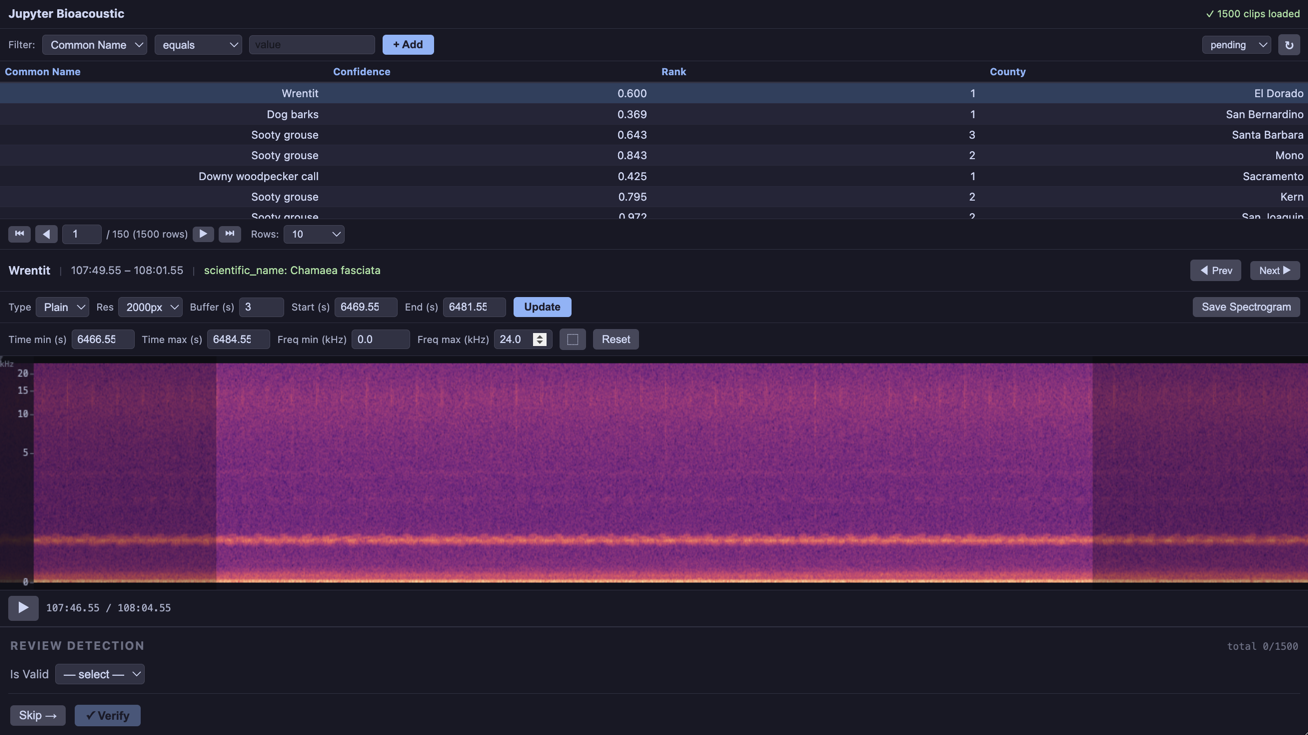
3.a Data Collection¶
file_config example
# config/forms/simple-examples.3a.yaml
select:
label: 'species'
column: 'common_name'
required: True
items:
path: CATEGORIES
value: 'common_name'BioacousticAnnotator(
data=DATA,
audio=AUDIO,
form_config='config/forms/simple-examples-3a.yaml'
).open()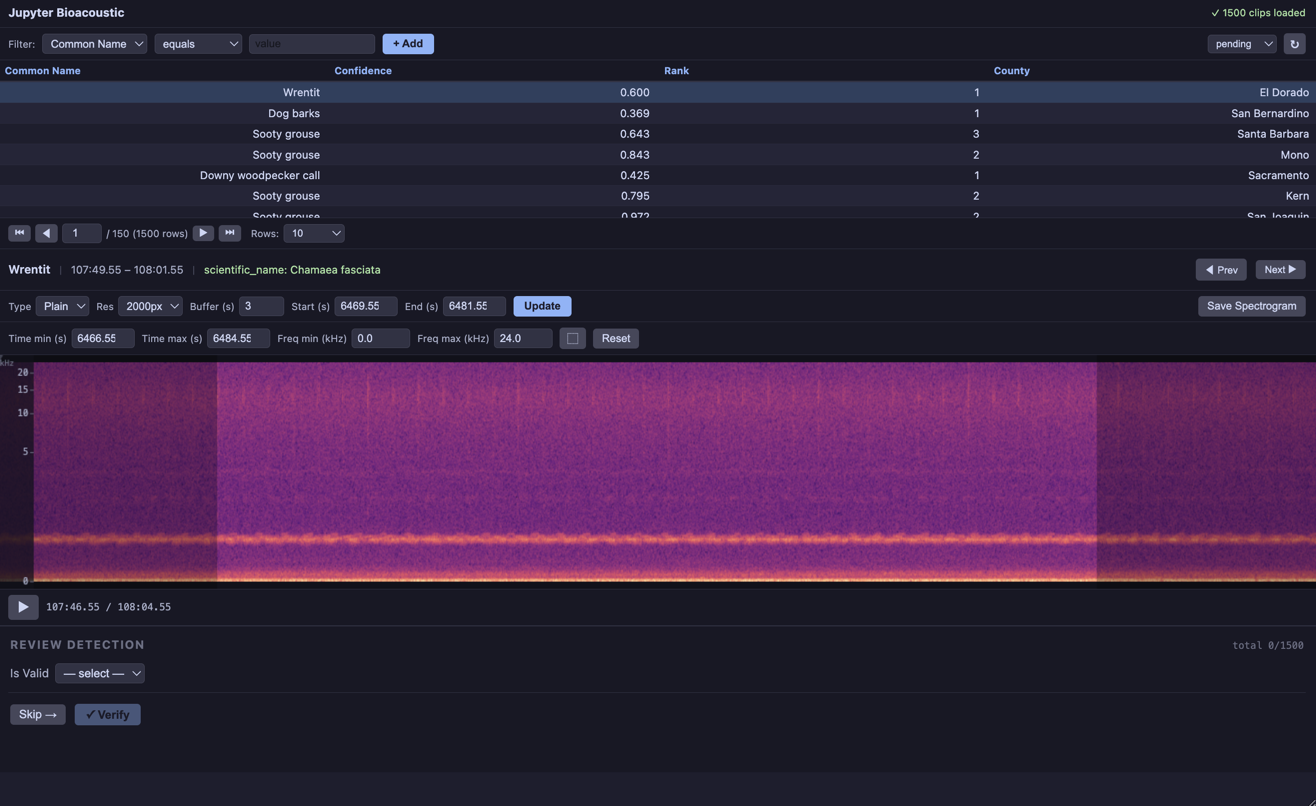
3.b Model Review¶
config example
The advantage to config over form_config is that you can pass the full range of parameters to BioacousticAnnotator. Note however, in this example we are passing only form_config. You can dive deeper into the full set of parameters here. We have kept data and audio as inline params, since this would allow a user to use the same config for multiple different data and audio values.
# config/simple-examples-3b.yaml
form_config:
select:
label: Is Valid
column: is_valid
required: true
items:
- value: 'yes'
- value: 'no'
form: correction_form
dynamic_forms:
correction_form:
- select:
label: corrected species
column: corrected_common_name
required: true
items:
path: "data/categories-small.csv"
value: "common_name"
required': true
filter_box': true
not_available': 'Unknown Species'BioacousticAnnotator(
data=DATA,
audio=AUDIO,
config='config/simple-examples-3c.yaml'
).open()