The most basic configuration of BioacousticAnnotator is a simple Player / Visualizer.
BioacousticAnnotator(data='detections.csv', audio='recording.flac').open()Here we have used a CSV for the source data. We could similarly have provided a Parquet, or JSONL file path, an API-endpoit, a Database Query (using DuckDB), or have passed a dataframe directly. For audio we have passed a path to an audio file. For this example all the clips will come from the same file. You can also pass an audio_column so that each clip can be read from different audio file. The audio files can be local paths or be read over https://, s3://, and gs://.
For more advanced configurations it is suggested that to use a configurtion file.
Here’s a basic example:
data_columns:
- 'common_name'
- 'confidence'
- 'rank'
- 'county'
ident_column: 'common_name'
display_columns: ['scientific_name']
capture: 'Save Spectrogram'
capture_dir: 'spectrograms'
form_config: 'config/forms/simple-examples-3c2.yaml'
We have not included the data and audio parameters. Although they could be included, by keeping them external to the config the user can use the same configuration files across projects and end-users.
FORM CONFIG
We could have included the form config directly, however seperating the form config allows different projects to use the same forms. Forms themeselves can range to a simple select-dropdown to multipart dynamic forms. Even the most complex forms however are easy to configure.
Here we create a “species” select list populated with values from a csv:

form:
- select:
label: 'species'
column: 'common_name'
required: true
items:
path: 'data/categories-small.csv'
value: 'common_name'
Additional functionality is easily added. The config below, again creates a “species” select list but now adds a filter box, an Unknown-option and the ability to add custom-values:
The following configuration below adds a custom title and a progress-tracker, passes the source id to the output dataset with name detection_id, and asks the user if the model has correctly identified the species.
title:
value: REVIEW DETECTION
progress_tracker: true
pass_value:
source_column: id
column: detection_id
form:
- select:
label: Is Valid
column: is_valid
required: true
items:
- label: 'yes'
value: 'yes'
form: confirmed_form
- label: 'no'
value: 'no'
form: rejected_form
dynamic_forms:
- confirmed_form:
- select:
label: confidence
column: reviewer_confidence
items: [low, medium, high]
- textbox:
label: notes
column: notes
- rejected_form:
- select:
label: corrected species
column: corrected_common_name
required: true
items:
path: data/categories-small.csv
value: common_name
filter_box: true
custom_value: true
not_available:
label: Unknown Species
value: unknown
- select:
label: rejection reason
column: rejection_reason
required: true
items:
- noise
- wrong species
- overlapping signals
- too faint
- other
- number:
label: signal quality (1-5)
column: signal_quality
min: 1
max: 5
step: 1
- checkbox:
label: flag for expert review
column: flagged
- textbox:
label: notes
column: notes
multiline: true
submission_buttons:
line: true
next:
label: Skip
submit:
label: Verify
If the response is yes the confirmed_form allows for the user to add additional detials.
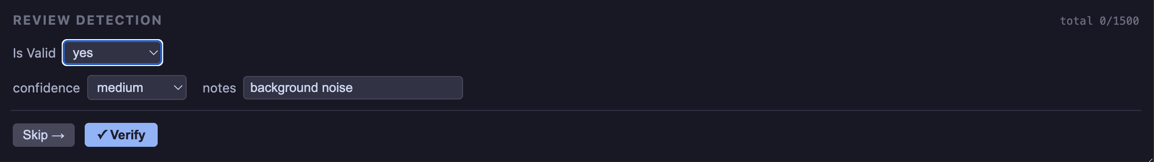
If the response is no the rejected_form allows the user to select the correct species, ask for a review, etc.
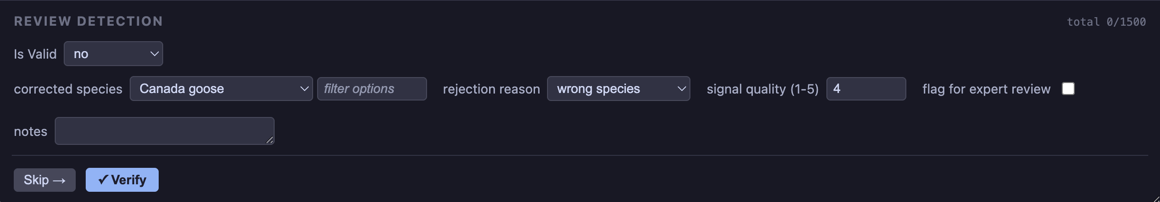
Additional examples are given in the Customizable Forms section.