from joshpy.registry import RunRegistry
# Load registry from disk (created by manual-workflow.qmd or sweep-manager.qmd)
REGISTRY_PATH = "demo_registry.duckdb"
registry = RunRegistry(REGISTRY_PATH)Analysis & Visualization
Introduction
This demo focuses on exploring and visualizing simulation results after they’ve been collected. The analysis layer in joshpy is decoupled from orchestration - it only needs a RunRegistry containing data. Whether the data came from SweepManager, manual workflow, or someone else’s experiment doesn’t matter.
Prerequisites: Run either Manual Workflow or SweepManager Workflow first to generate demo_registry.duckdb.
For orchestration workflows, see:
- Manual Workflow - Step-by-step control using individual components
- SweepManager Workflow - Simplified workflow with
SweepManager
This demo covers:
- Data Discovery - What’s in the registry?
- Diagnostic Plots - Quick matplotlib visualizations
- Custom Queries -
DiagnosticQueriesfor common patterns - Direct SQL - Full access to DuckDB for advanced analysis
- R/ggplot2 - Publication-quality figures
Setup: Load Registry
Load the registry saved by a previous orchestration demo:
Part 1: Data Discovery
Before plotting, discover what data is available in the registry. This step is essential - column names come from your Josh exports, not from joshpy.
Always run list_export_variables() and list_config_parameters() before writing queries. See Best Practices: Schema & Querying for common query patterns and aggregation guidance.
Full Summary
summary = registry.get_data_summary()
print(summary)Registry Data Summary
========================================
Sessions: 1
Configs: 10
Runs: 10
Rows: 43,890
Variables: averageAge, averageHeight
Entity types: patch
Parameters: maxGrowth
Steps: 0 - 10
Replicates: 0 - 2
Spatial extent: lon [-115.37, -114.40], lat [33.41, 33.68]Specific Queries
# What variables were exported from the simulation?
registry.list_export_variables()['averageAge', 'averageHeight']# What parameters were swept?
registry.list_config_parameters()['maxGrowth']# What entity types are in the data?
registry.list_entity_types()['patch']Variable Columns
# List all variable columns in the cell_data table
registry.list_variable_columns()['averageAge', 'averageHeight']Part 2: Diagnostic Plots (SimulationDiagnostics)
The SimulationDiagnostics class provides quick matplotlib-based visualizations for simulation sanity checks. These are designed for rapid exploration, not publication.
from joshpy.diagnostics import SimulationDiagnostics
diag = SimulationDiagnostics(registry)Time Series
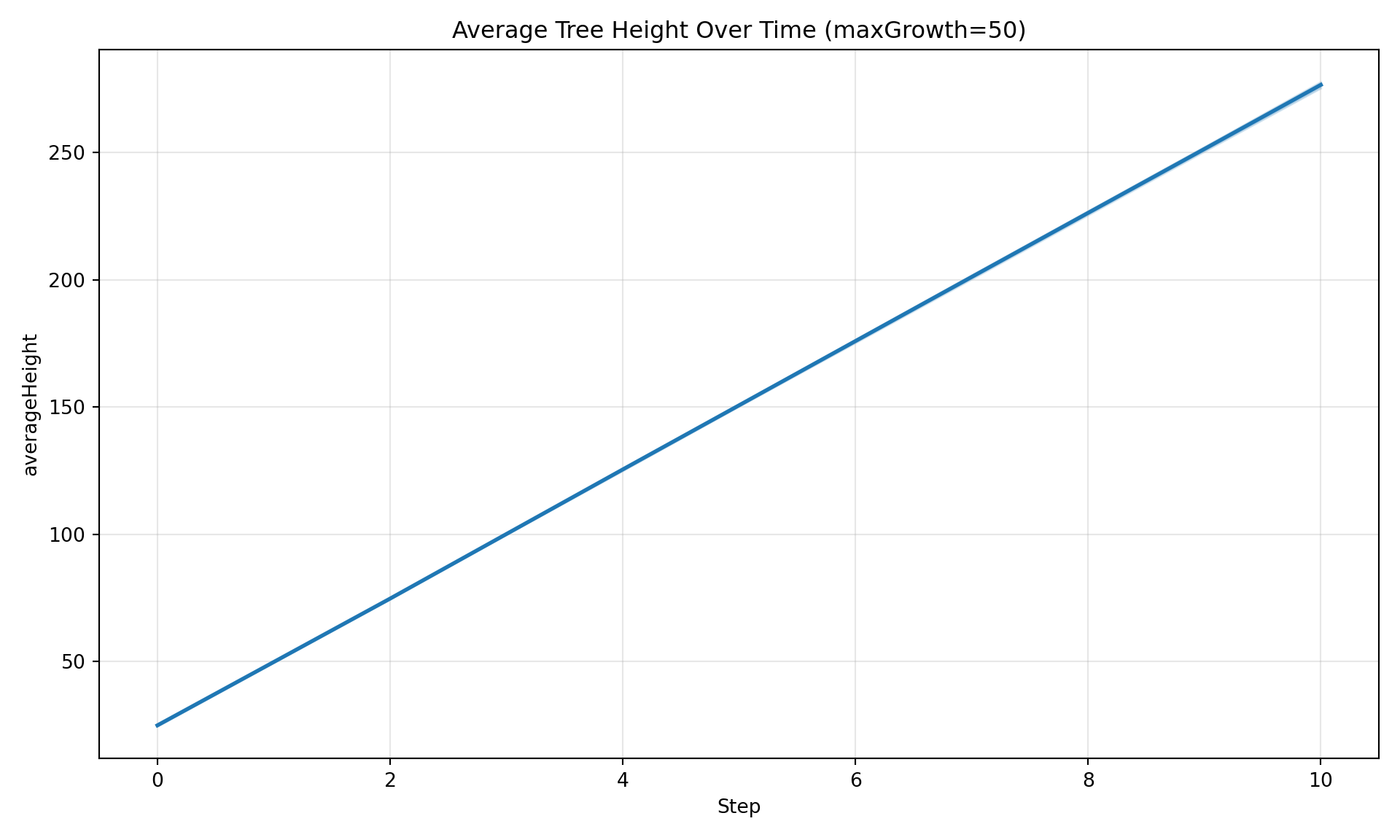
Plot how a variable evolves over simulation steps. By default, spatially aggregates across patches and shows uncertainty bands across replicates.
# Filter by parameter value
diag.plot_timeseries(
"averageHeight",
maxGrowth=50,
title="Average Tree Height Over Time (maxGrowth=50)",
)
Aggregation Options
# Sum across patches
diag.plot_timeseries("averageHeight", maxGrowth=50, aggregate="sum", title="Sum")
# Min across patches
diag.plot_timeseries("averageHeight", maxGrowth=50, aggregate="min", title="Min")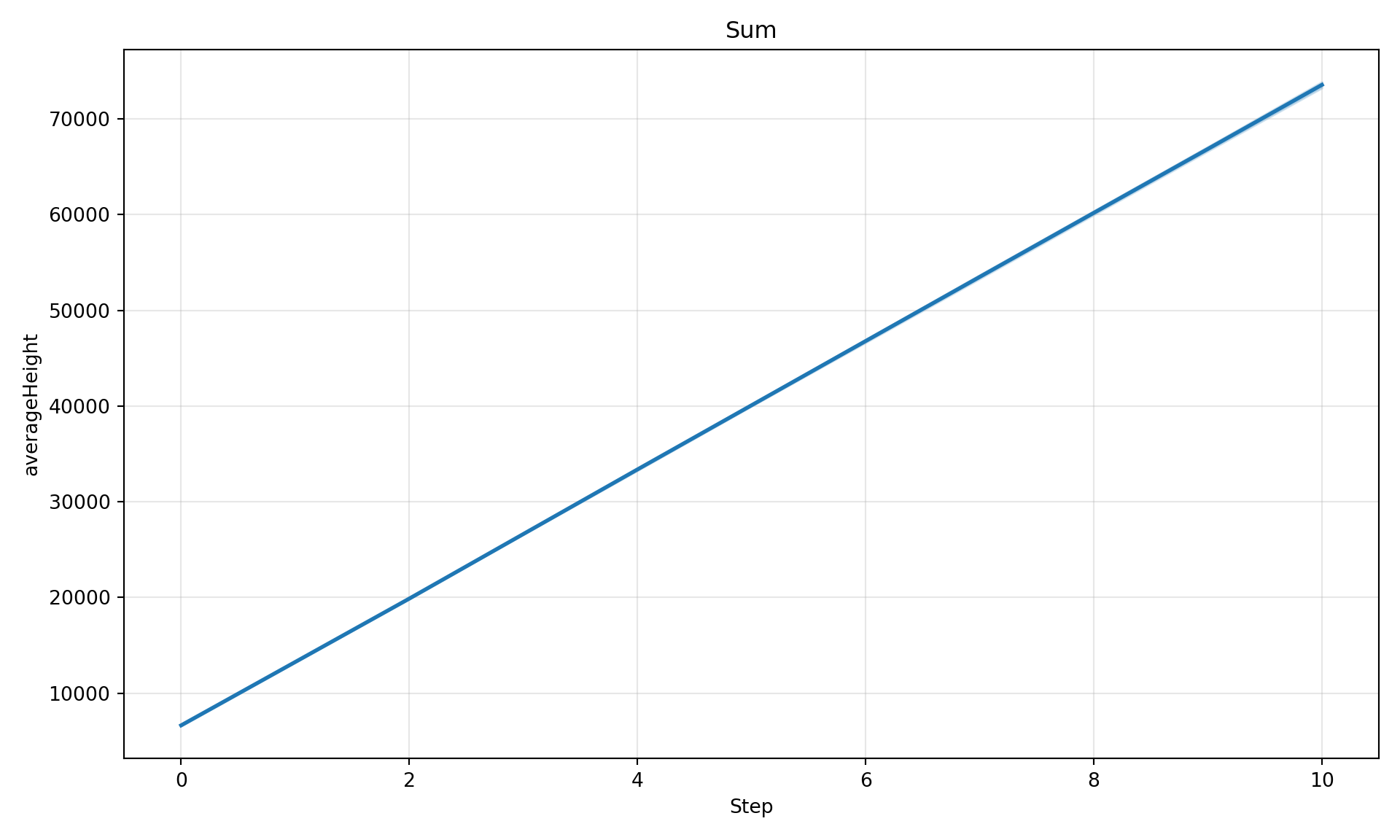
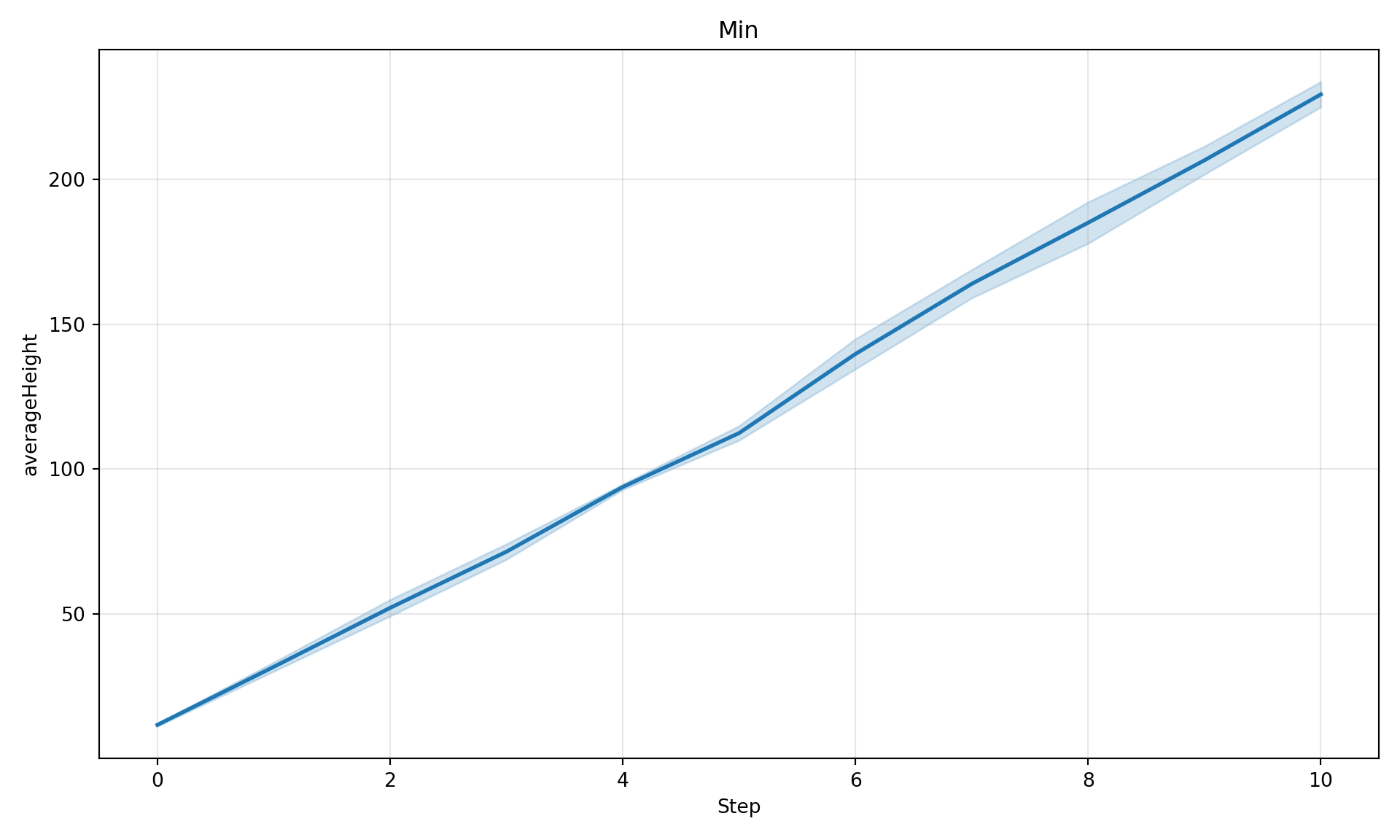
Parameter Comparison
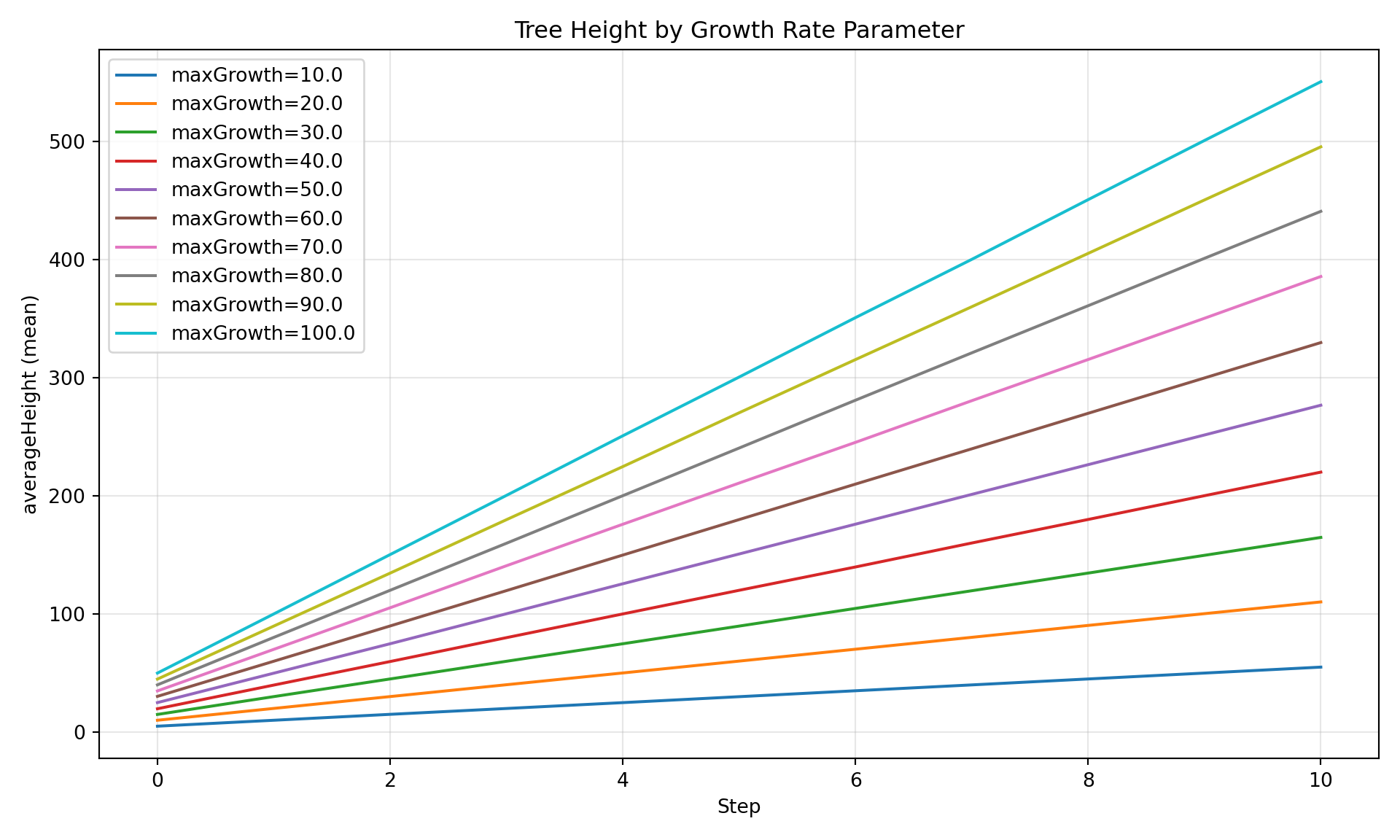
Compare a variable across different parameter values - ideal for sweep results.
diag.plot_comparison(
"averageHeight",
group_by="maxGrowth",
title="Tree Height by Growth Rate Parameter",
)
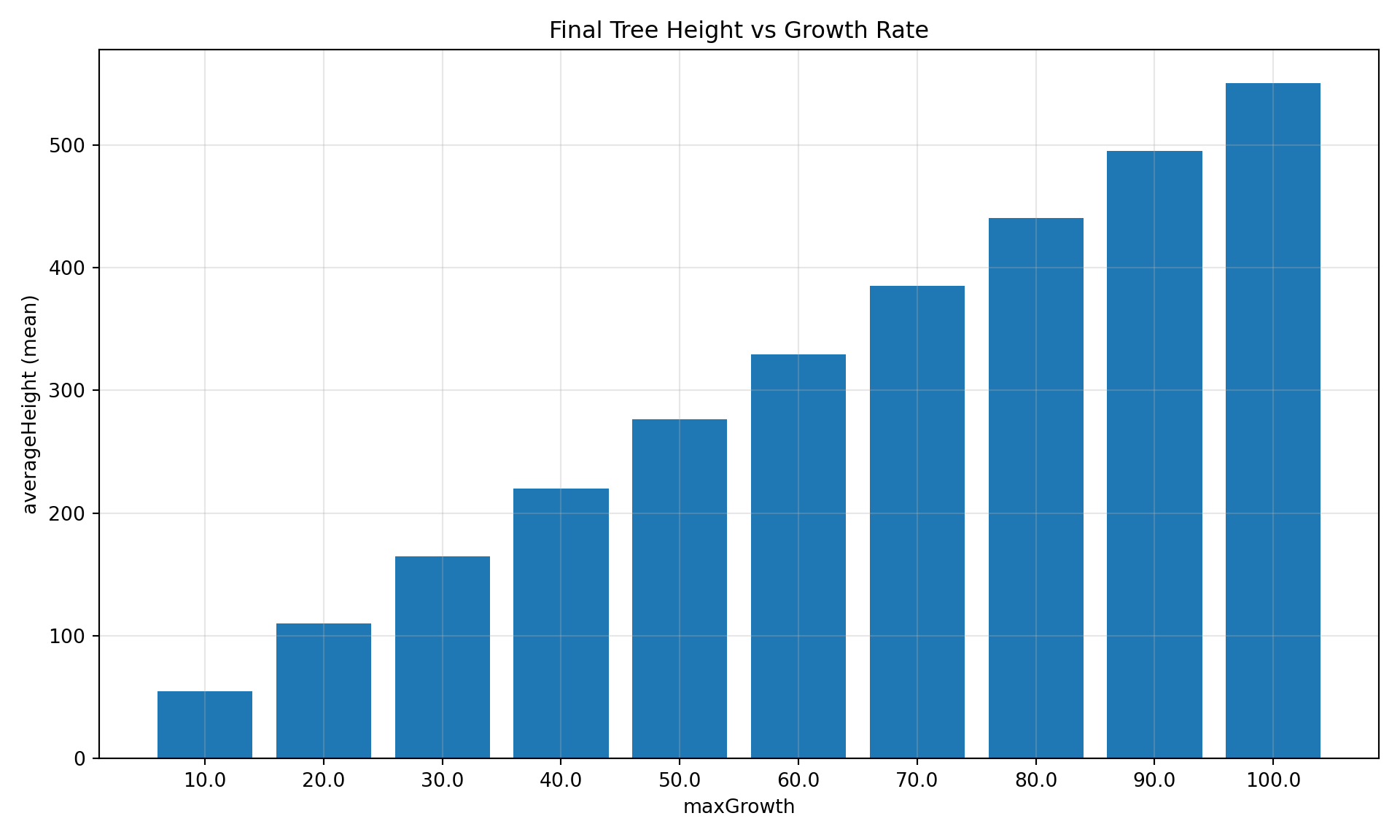
Bar Chart at Specific Step
diag.plot_comparison(
"averageHeight",
group_by="maxGrowth",
step=10,
title="Final Tree Height vs Growth Rate",
)
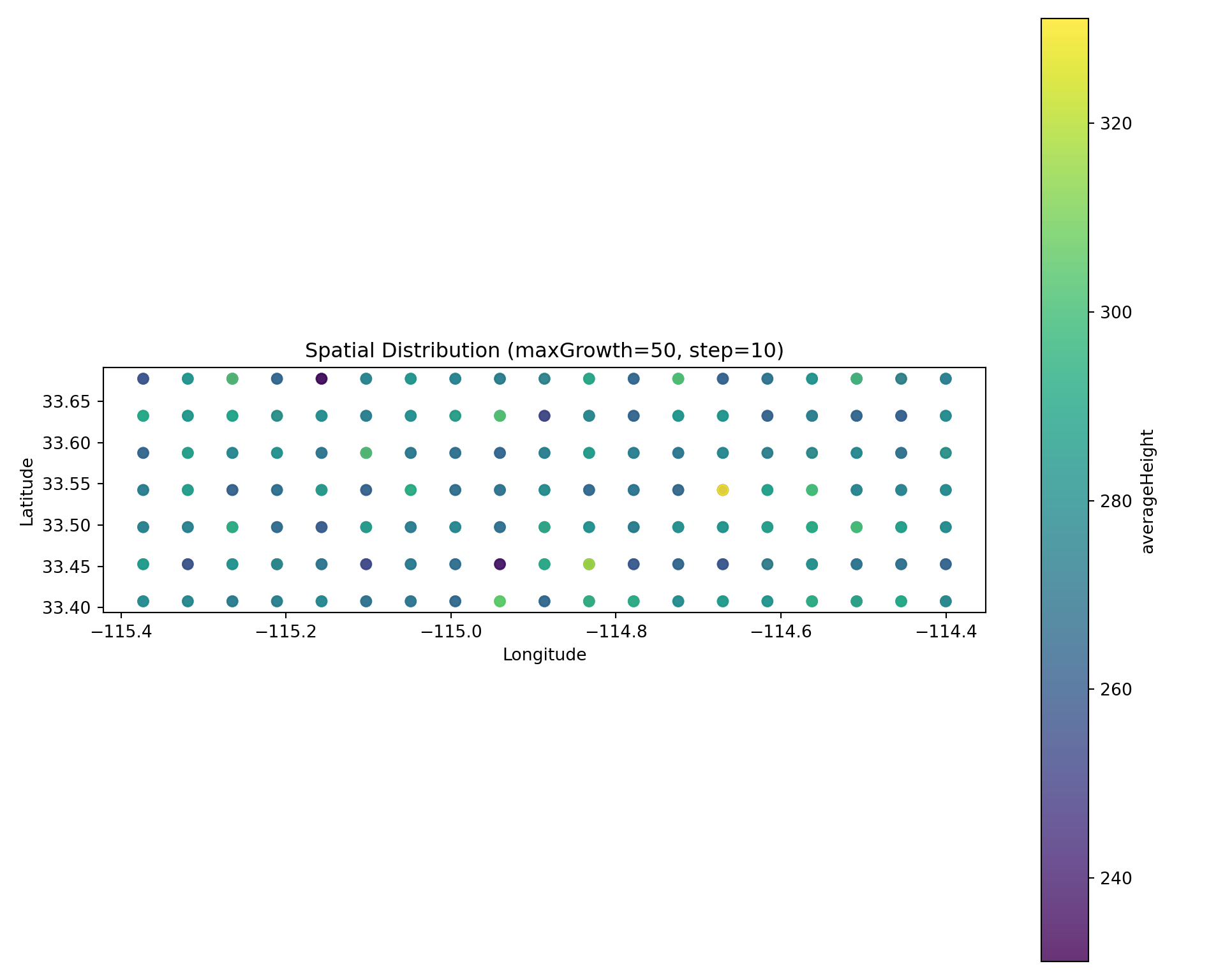
Spatial Snapshot
Scatter plot colored by value at a specific timestep:
# Get a run hash for maxGrowth=50
run_hash_50 = registry.get_runs_by_parameters(maxGrowth=50)[0]["run_hash"]
diag.plot_spatial(
"averageHeight",
step=10,
run_hash=run_hash_50,
title="Spatial Distribution (maxGrowth=50, step=10)",
)
Saving Figures
All plot methods return a matplotlib Figure that can be saved:
fig = diag.plot_comparison(
"averageHeight",
group_by="maxGrowth",
show=False, # Don't display inline
)
fig.savefig("/tmp/comparison_plot.png", dpi=150, bbox_inches="tight")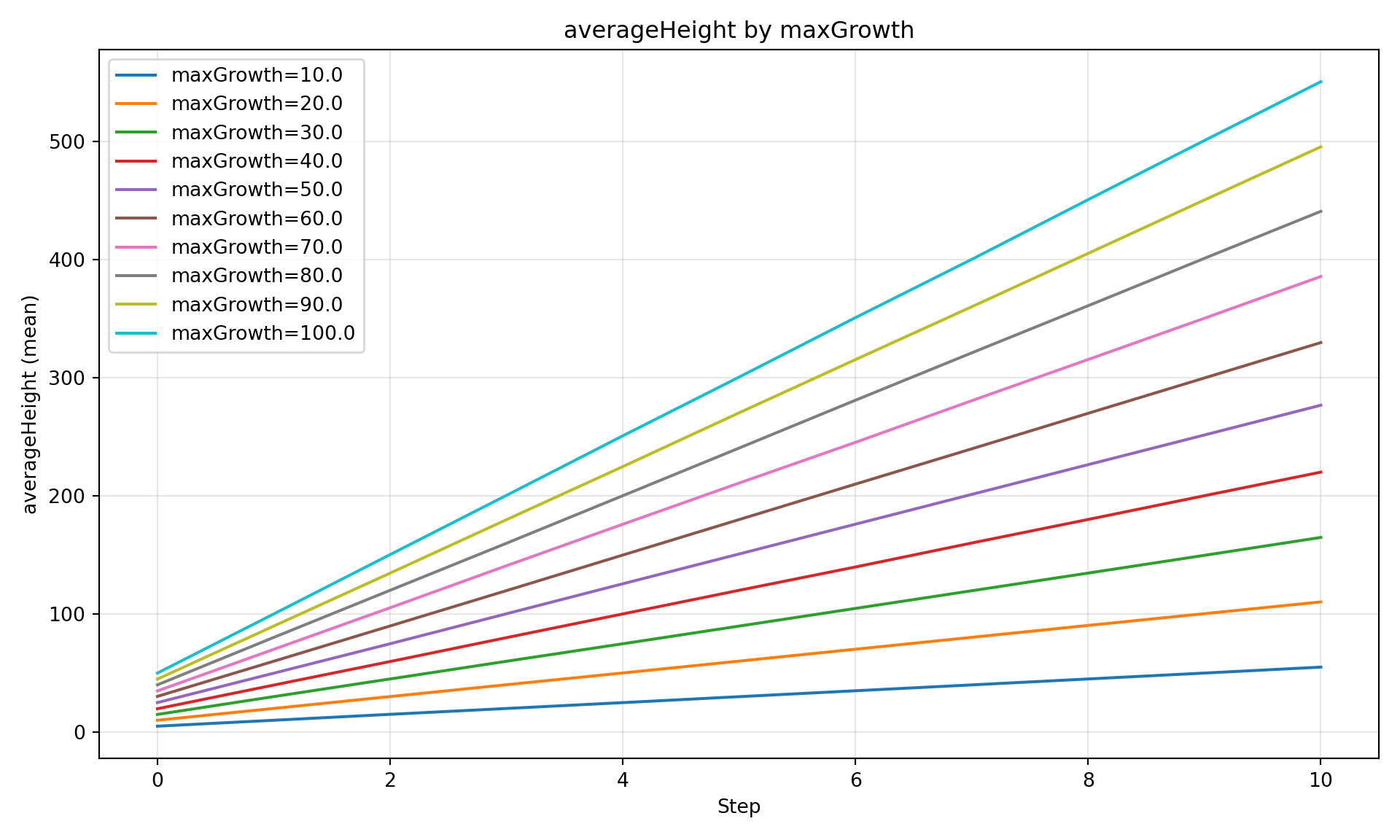
Part 3: Custom Queries (DiagnosticQueries)
For more control than SimulationDiagnostics, use DiagnosticQueries directly. These return pandas DataFrames for further processing.
from joshpy.cell_data import DiagnosticQueries
queries = DiagnosticQueries(registry)Parameter Comparison
df = queries.get_parameter_comparison(
variable="averageHeight",
param_name="maxGrowth",
)
df.head(15)| param_value | step | mean_value | std_value | n_cells | |
|---|---|---|---|---|---|
| 0 | 10.0 | 0 | 4.990825 | 0.914275 | 399 |
| 1 | 10.0 | 1 | 10.054546 | 1.244391 | 399 |
| 2 | 10.0 | 2 | 15.012953 | 1.548240 | 399 |
| 3 | 10.0 | 3 | 20.005701 | 1.847471 | 399 |
| 4 | 10.0 | 4 | 25.058594 | 2.014672 | 399 |
| 5 | 10.0 | 5 | 30.031544 | 2.194709 | 399 |
| 6 | 10.0 | 6 | 35.077375 | 2.397354 | 399 |
| 7 | 10.0 | 7 | 40.056394 | 2.636158 | 399 |
| 8 | 10.0 | 8 | 45.132441 | 2.798784 | 399 |
| 9 | 10.0 | 9 | 50.117931 | 2.962958 | 399 |
| 10 | 10.0 | 10 | 55.069338 | 3.150577 | 399 |
| 11 | 20.0 | 0 | 9.771987 | 1.809808 | 399 |
| 12 | 20.0 | 1 | 19.800545 | 2.610955 | 399 |
| 13 | 20.0 | 2 | 29.743859 | 3.267051 | 399 |
| 14 | 20.0 | 3 | 39.724835 | 3.775138 | 399 |
Replicate Uncertainty
Get statistics across replicates for a specific run:
df = queries.get_replicate_uncertainty(
variable="averageHeight",
run_hash=run_hash_50,
)
df.head(10)| step | mean | std | ci_low | ci_high | n_replicates | |
|---|---|---|---|---|---|---|
| 0 | 0 | 25.044947 | 0.473936 | 24.508648 | 25.581246 | 3 |
| 1 | 1 | 50.248364 | 0.317795 | 49.888752 | 50.607976 | 3 |
| 2 | 2 | 75.673786 | 0.321899 | 75.309530 | 76.038042 | 3 |
| 3 | 3 | 100.530033 | 0.484435 | 99.981854 | 101.078212 | 3 |
| 4 | 4 | 125.526812 | 0.348489 | 125.132467 | 125.921157 | 3 |
| 5 | 5 | 150.787472 | 0.325004 | 150.419702 | 151.155242 | 3 |
| 6 | 6 | 175.415496 | 0.752038 | 174.564501 | 176.266491 | 3 |
| 7 | 7 | 200.361078 | 0.728033 | 199.537246 | 201.184910 | 3 |
| 8 | 8 | 225.421147 | 0.736903 | 224.587278 | 226.255017 | 3 |
| 9 | 9 | 250.298304 | 0.804500 | 249.387943 | 251.208665 | 3 |
Spatial Snapshot
Get all spatial data for a given timestep:
df = queries.get_spatial_snapshot(
step=10,
variable="averageHeight",
run_hash=run_hash_50,
)
df.head(10)| longitude | latitude | value | |
|---|---|---|---|
| 0 | -114.508196 | 33.452687 | 269.959912 |
| 1 | -115.210829 | 33.497653 | 287.364517 |
| 2 | -115.102732 | 33.452687 | 276.273301 |
| 3 | -114.778439 | 33.632551 | 272.278324 |
| 4 | -114.778439 | 33.587585 | 271.014116 |
| 5 | -115.318927 | 33.542619 | 276.615131 |
| 6 | -114.562244 | 33.407720 | 268.089078 |
| 7 | -114.508196 | 33.542619 | 276.041247 |
| 8 | -114.724391 | 33.542619 | 255.923528 |
| 9 | -115.210829 | 33.587585 | 309.713661 |
Get All Variables at Once
Retrieve all exported variables for a specific step:
df = queries.get_all_variables_at_step(
step=10,
run_hash=run_hash_50,
)
df.head(5)| longitude | latitude | position_x | position_y | averageAge | averageHeight | |
|---|---|---|---|---|---|---|
| 0 | -114.508196 | 33.452687 | 16.5 | 5.5 | 11.0 | 269.959912 |
| 1 | -115.210829 | 33.497653 | 3.5 | 4.5 | 11.0 | 287.364517 |
| 2 | -115.102732 | 33.452687 | 5.5 | 5.5 | 11.0 | 276.273301 |
| 3 | -114.778439 | 33.632551 | 11.5 | 1.5 | 11.0 | 272.278324 |
| 4 | -114.778439 | 33.587585 | 11.5 | 2.5 | 11.0 | 271.014116 |
Part 4: Direct SQL Queries
For full flexibility, query DuckDB directly.
Simple Aggregations
# Mean, min, max across all cells per step
result = registry.query("""
SELECT
step,
AVG(averageHeight) as mean_height,
MIN(averageHeight) as min_height,
MAX(averageHeight) as max_height,
STDDEV(averageHeight) as std_height,
COUNT(*) as n_cells
FROM cell_data
GROUP BY step
ORDER BY step
""")
result.df()| step | mean_height | min_height | max_height | std_height | n_cells | |
|---|---|---|---|---|---|---|
| 0 | 0 | 27.681788 | 2.821725 | 78.752073 | 15.629399 | 3990 |
| 1 | 1 | 55.242309 | 6.998114 | 136.550564 | 29.959751 | 3990 |
| 2 | 2 | 82.821625 | 10.546986 | 197.042799 | 44.345770 | 3990 |
| 3 | 3 | 110.122145 | 15.145431 | 246.139270 | 58.432869 | 3990 |
| 4 | 4 | 137.700887 | 18.950683 | 308.720026 | 72.853441 | 3990 |
| 5 | 5 | 165.319256 | 23.451497 | 359.712621 | 87.244514 | 3990 |
| 6 | 6 | 192.829216 | 27.630700 | 418.271207 | 101.571358 | 3990 |
| 7 | 7 | 220.272909 | 29.934035 | 479.443895 | 115.935966 | 3990 |
| 8 | 8 | 247.873201 | 33.037224 | 530.481536 | 130.335174 | 3990 |
| 9 | 9 | 275.381847 | 37.641517 | 587.912941 | 144.757198 | 3990 |
| 10 | 10 | 302.720721 | 42.661982 | 644.653712 | 159.088170 | 3990 |
Filtering on Variable Values
# Find cells where trees grew above a threshold
result = registry.query("""
SELECT
step,
COUNT(*) as n_cells_above_50,
AVG(averageHeight) as mean_height_of_those
FROM cell_data
WHERE averageHeight > 50
GROUP BY step
ORDER BY step
""")
result.df()| step | n_cells_above_50 | mean_height_of_those | |
|---|---|---|---|
| 0 | 0 | 365 | 56.565190 |
| 1 | 1 | 2163 | 78.597847 |
| 2 | 2 | 2841 | 104.594483 |
| 3 | 3 | 3179 | 130.525999 |
| 4 | 4 | 3385 | 156.514972 |
| 5 | 5 | 3585 | 180.574037 |
| 6 | 6 | 3591 | 210.357199 |
| 7 | 7 | 3591 | 240.296967 |
| 8 | 8 | 3610 | 269.244289 |
| 9 | 9 | 3802 | 286.643922 |
| 10 | 10 | 3971 | 303.936274 |
Percentile Analysis
# Get distribution statistics per step
result = registry.query("""
SELECT
step,
APPROX_QUANTILE(averageHeight, 0.25) as p25,
APPROX_QUANTILE(averageHeight, 0.50) as median,
APPROX_QUANTILE(averageHeight, 0.75) as p75,
APPROX_QUANTILE(averageHeight, 0.95) as p95
FROM cell_data
GROUP BY step
ORDER BY step
""")
result.df()| step | p25 | median | p75 | p95 | |
|---|---|---|---|---|---|
| 0 | 0 | 14.603601 | 26.594013 | 39.414013 | 54.510150 |
| 1 | 1 | 29.880371 | 54.580641 | 79.148126 | 104.793557 |
| 2 | 2 | 45.072850 | 82.426778 | 119.524511 | 153.619784 |
| 3 | 3 | 60.105815 | 109.673034 | 159.243393 | 202.819378 |
| 4 | 4 | 74.816438 | 136.922768 | 198.987215 | 252.059392 |
| 5 | 5 | 90.171734 | 164.280947 | 239.519276 | 301.707617 |
| 6 | 6 | 105.229610 | 191.610866 | 280.087840 | 350.872525 |
| 7 | 7 | 119.829756 | 219.377950 | 320.230818 | 400.050208 |
| 8 | 8 | 135.308450 | 246.712727 | 359.629457 | 450.522048 |
| 9 | 9 | 150.404325 | 273.748778 | 400.159652 | 500.730780 |
| 10 | 10 | 164.853972 | 300.675263 | 440.601935 | 550.360618 |
Joining with Config Parameters
Config parameters are stored as typed columns in the config_parameters table:
# Compare results across different parameter values
result = registry.query("""
SELECT
cp.maxGrowth,
cd.step,
AVG(cd.averageHeight) as mean_height,
STDDEV(cd.averageHeight) as std_height
FROM cell_data cd
JOIN config_parameters cp ON cd.run_hash = cp.run_hash
GROUP BY cp.maxGrowth, cd.step
ORDER BY cp.maxGrowth, cd.step
""")
result.df().head(15)| maxGrowth | step | mean_height | std_height | |
|---|---|---|---|---|
| 0 | 10.0 | 0 | 4.990825 | 0.914275 |
| 1 | 10.0 | 1 | 10.054546 | 1.244391 |
| 2 | 10.0 | 2 | 15.012953 | 1.548240 |
| 3 | 10.0 | 3 | 20.005701 | 1.847471 |
| 4 | 10.0 | 4 | 25.058594 | 2.014672 |
| 5 | 10.0 | 5 | 30.031544 | 2.194709 |
| 6 | 10.0 | 6 | 35.077375 | 2.397354 |
| 7 | 10.0 | 7 | 40.056394 | 2.636158 |
| 8 | 10.0 | 8 | 45.132441 | 2.798784 |
| 9 | 10.0 | 9 | 50.117931 | 2.962958 |
| 10 | 10.0 | 10 | 55.069338 | 3.150577 |
| 11 | 20.0 | 0 | 9.771987 | 1.809808 |
| 12 | 20.0 | 1 | 19.800545 | 2.610955 |
| 13 | 20.0 | 2 | 29.743859 | 3.267051 |
| 14 | 20.0 | 3 | 39.724835 | 3.775138 |
# Filter by parameter value (maxGrowth > 50)
result = registry.query("""
SELECT
cd.step,
AVG(cd.averageHeight) as mean_height
FROM cell_data cd
JOIN config_parameters cp ON cd.run_hash = cp.run_hash
WHERE cp.maxGrowth > 50
GROUP BY cd.step
ORDER BY cd.step
""")
result.df()| step | mean_height | |
|---|---|---|
| 0 | 0 | 40.272835 |
| 1 | 1 | 80.377612 |
| 2 | 2 | 120.359578 |
| 3 | 3 | 159.985498 |
| 4 | 4 | 200.152581 |
| 5 | 5 | 240.329104 |
| 6 | 6 | 280.418718 |
| 7 | 7 | 320.387910 |
| 8 | 8 | 360.484870 |
| 9 | 9 | 400.502498 |
| 10 | 10 | 440.331969 |
Export to File
# Export query results to CSV
registry.query("""
SELECT step, AVG(averageHeight) as mean_height
FROM cell_data
GROUP BY step
ORDER BY step
""").df().to_csv("/tmp/height_by_step.csv", index=False)Part 5: R/ggplot2 Visualization
For publication-quality figures, there are three routes to get data from joshpy into R:
- CSV Export - Simple, always works
- Direct DuckDB - Custom SQL queries from R, no Python needed
py$dfvia reticulate - Zero-copy Python/R interop
Route 1: Export to CSV (Simple, Always Works)
# Python exports to CSV
csv_df = queries.get_parameter_comparison(
variable="averageHeight",
param_name="maxGrowth",
)
csv_df.to_csv("/tmp/sweep_results.csv", index=False)# R reads CSV
df_csv <- read.csv("/tmp/sweep_results.csv")
df_csv$param_value <- as.numeric(df_csv$param_value)
head(df_csv, 5) param_value step mean_value std_value n_cells
1 10 0 4.990825 0.9142747 399
2 10 1 10.054546 1.2443914 399
3 10 2 15.012953 1.5482401 399
4 10 3 20.005701 1.8474706 399
5 10 4 25.058594 2.0146724 399Pros: Always works, simple, portable
Cons: File I/O overhead, temp file management
Route 2: Direct DuckDB from R (Custom Queries)
R can query DuckDB directly with full SQL flexibility:
library(DBI)
library(duckdb)
con <- dbConnect(duckdb(), "demo_registry.duckdb", read_only = TRUE)
df_duckdb <- dbGetQuery(con, "
SELECT
cp.maxGrowth as param_value,
cd.step,
AVG(cd.averageHeight) as mean_value,
STDDEV(cd.averageHeight) as std_value
FROM cell_data cd
JOIN config_parameters cp ON cd.run_hash = cp.run_hash
GROUP BY cp.maxGrowth, cd.step
ORDER BY cp.maxGrowth, cd.step
")
dbDisconnect(con, shutdown = TRUE)
head(df_duckdb, 10) param_value step mean_value std_value
1 10 0 4.990825 0.9142747
2 10 1 10.054546 1.2443914
3 10 2 15.012953 1.5482401
4 10 3 20.005701 1.8474706
5 10 4 25.058594 2.0146724
6 10 5 30.031544 2.1947085
7 10 6 35.077375 2.3973544
8 10 7 40.056394 2.6361577
9 10 8 45.132441 2.7987839
10 10 9 50.117931 2.9629578Pros: Full SQL flexibility, no Python needed for custom queries
Cons: Requires r-duckdb package (linux-64 only in pixi)
Route 3: py$df via reticulate (Zero-Copy)
In theory, you can access Python DataFrames directly via py$df:
# This should work:
# df <- reticulate::py_to_r(py$sweep_df)We’re working on this - for now, Routes 1 and 2 are recommended.
Plotting with ggplot2
The following examples use df_duckdb from Route 2 (direct DuckDB query). We assign it to df for cleaner ggplot code:
# Use the DuckDB query result for plotting
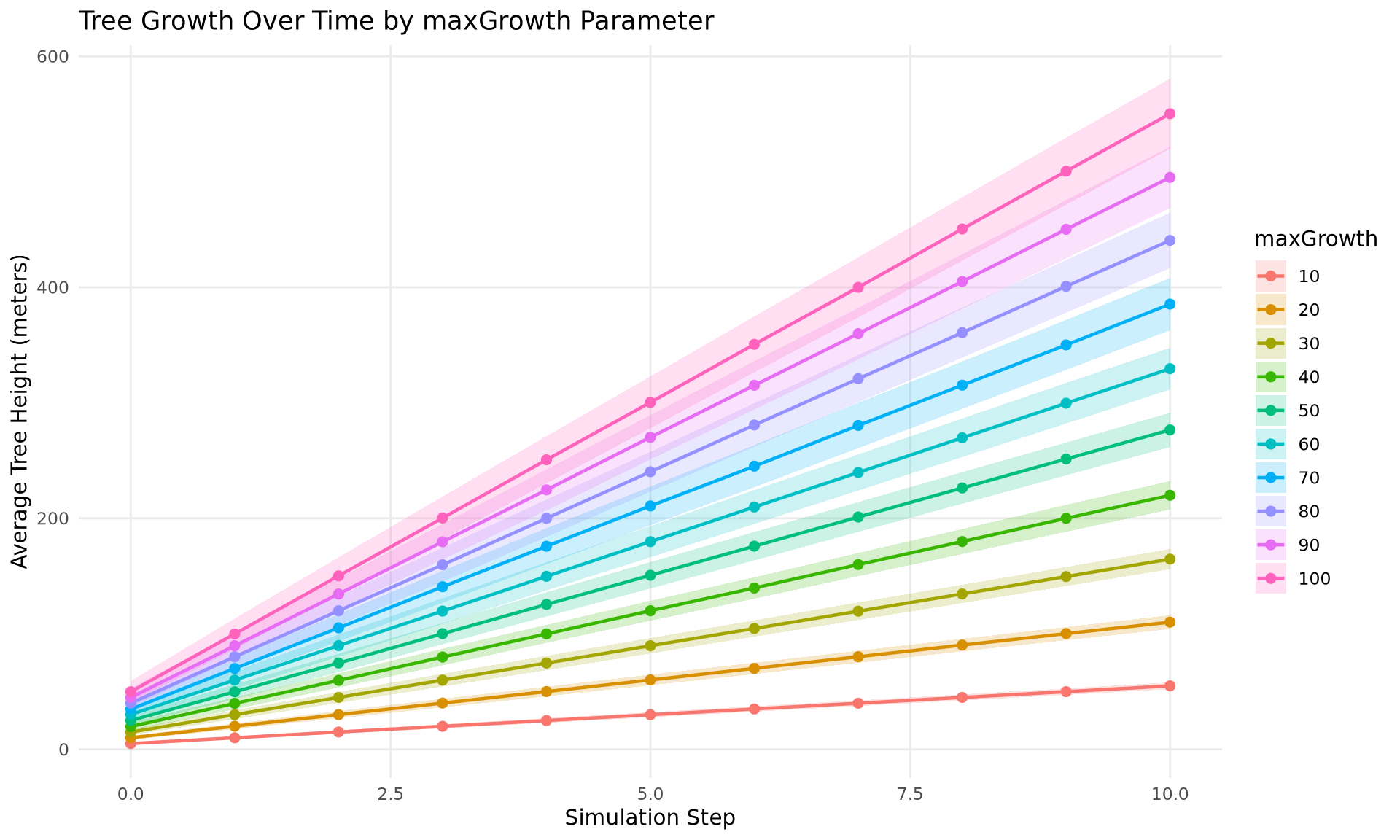
df <- df_duckdbTime Series with Uncertainty Ribbon
ggplot(df, aes(x = step, y = mean_value,
color = factor(param_value),
fill = factor(param_value))) +
geom_ribbon(aes(ymin = mean_value - std_value,
ymax = mean_value + std_value),
alpha = 0.2, color = NA) +
geom_line(linewidth = 0.8) +
geom_point(size = 2) +
labs(
x = "Simulation Step",
y = "Average Tree Height (meters)",
title = "Tree Growth Over Time by maxGrowth Parameter",
color = "maxGrowth",
fill = "maxGrowth"
) +
theme_minimal() +
theme(
legend.position = "right",
panel.grid.minor = element_blank()
)
Bar Chart with Error Bars
final_df <- df |>
dplyr::filter(step == max(step))
ggplot(final_df, aes(x = param_value, y = mean_value)) +
geom_col(fill = "steelblue", alpha = 0.7) +
geom_errorbar(
aes(ymin = mean_value - std_value, ymax = mean_value + std_value),
width = 3,
linewidth = 0.6
) +
labs(
x = "maxGrowth (meters/step)",
y = "Final Average Height (meters)",
title = "Final Tree Height vs Growth Rate"
) +
theme_minimal() +
theme(panel.grid.minor = element_blank())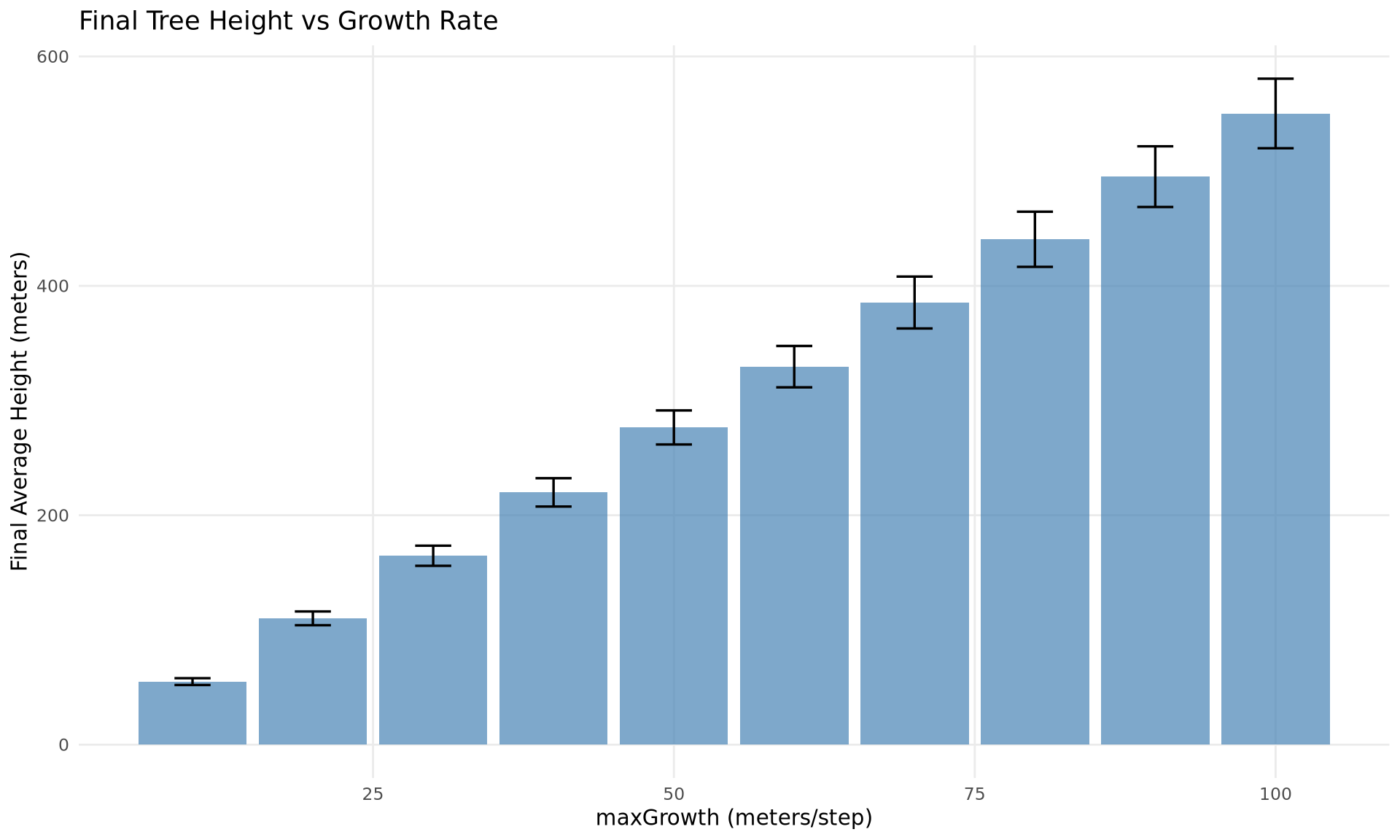
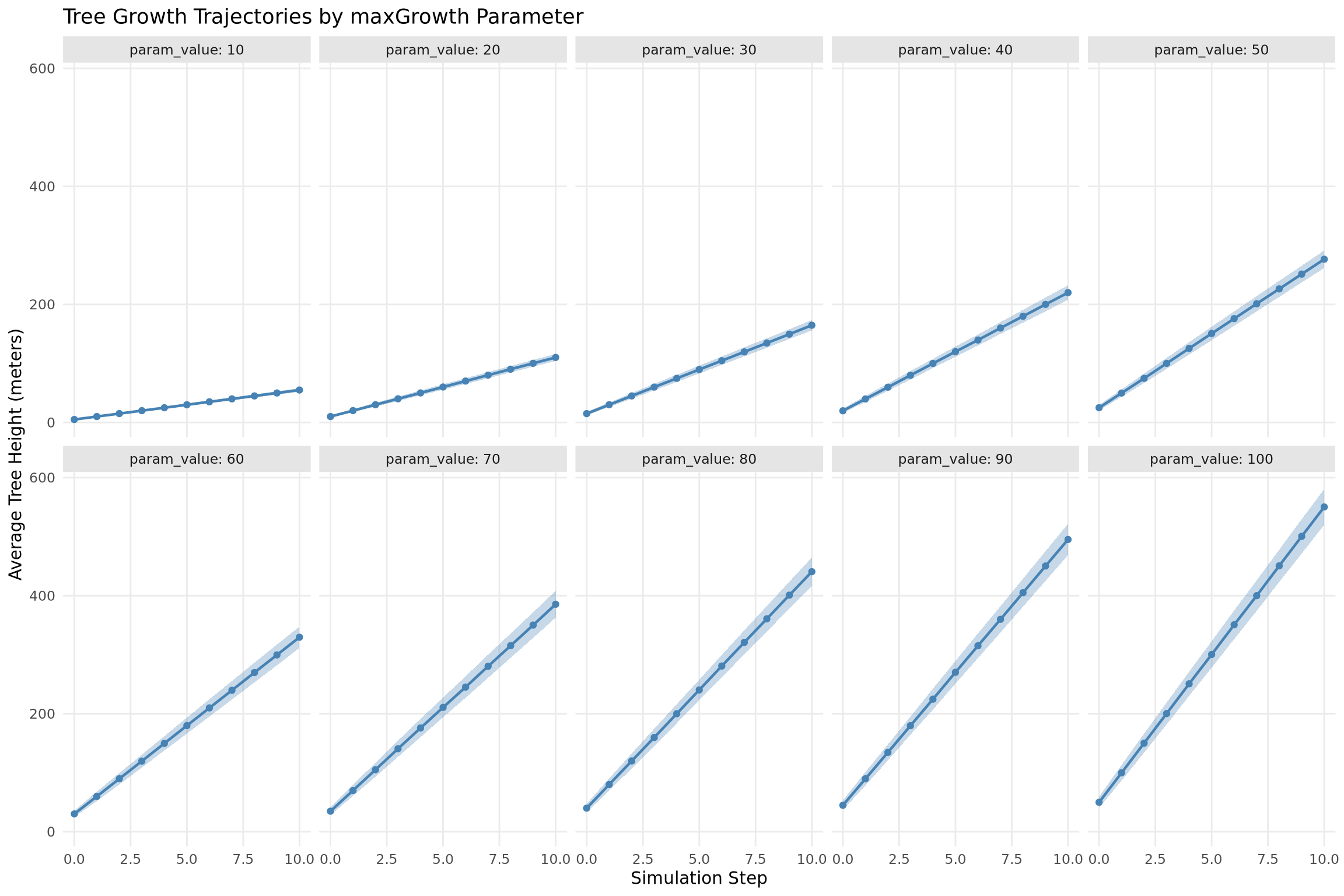
Faceted Plot
ggplot(df, aes(x = step, y = mean_value)) +
geom_ribbon(aes(ymin = mean_value - std_value,
ymax = mean_value + std_value),
alpha = 0.3, fill = "steelblue") +
geom_line(color = "steelblue", linewidth = 0.8) +
geom_point(color = "steelblue", size = 1.5) +
facet_wrap(~param_value, ncol = 5, labeller = label_both) +
labs(
x = "Simulation Step",
y = "Average Tree Height (meters)",
title = "Tree Growth Trajectories by maxGrowth Parameter"
) +
theme_minimal() +
theme(
panel.grid.minor = element_blank(),
strip.background = element_rect(fill = "gray90", color = NA)
)
Summary
This demo covered the joshpy analysis workflow:
| Task | Tool | Notes |
|---|---|---|
| Discovery | registry.get_data_summary() |
What’s in the registry? |
| Quick plots | SimulationDiagnostics |
matplotlib-based, fast iteration |
| DataFrames | DiagnosticQueries |
Returns pandas for further processing |
| Full SQL | registry.query() |
Direct DuckDB access |
| R (CSV) | read.csv() |
Simple, always works |
| R (DuckDB) | dbConnect(duckdb(), ...) |
Custom SQL directly from R |
| R (reticulate) | py$df |
Zero-copy Python/R interop |
Key Points:
- Analysis is decoupled from orchestration - same workflow for any data source
- Export variables are typed columns in
cell_data - Config parameters are typed columns in
config_parameters - R can query DuckDB directly
- Three routes for R: CSV export, direct DuckDB, or
py$dfvia reticulate
Cleanup
registry.close()